By ChatGPT, Claude, DeepSeek, Gemini, Grok, with W.H.L.
W.H.L.: Hi There! Could you provide some info about the latest UIUC paper in Cell: what is novel, why it is a significant breakthrough?
ChatGPT: UIUC’s 2026 Cell Paper — “Bringing the genetically minimal cell to life on a computer in 4D”



A research team at the University of Illinois Urbana–Champaign (UIUC) led by Zan Luthey-Schulten recently published a major paper in Cell describing the first full spatial-temporal simulation of a living minimal cell through an entire life cycle. (chemistry.illinois.edu)
The study models the bacterium JCVI-syn3A, a genetically minimal organism engineered by the J. Craig Venter Institute that contains only the genes necessary for life. (chemistry.illinois.edu)
Below is what’s new and why scientists consider it a breakthrough.
What the Paper Actually Did
1. Simulated a whole living cell over time
The researchers built a 3-dimensional, time-evolving (“4D”) computational model that tracks essentially every molecule inside a cell during its entire cell cycle (~100 minutes). (Cell)
The simulation includes:
- DNA replication
- RNA transcription
- protein translation
- metabolism
- spatial molecular motion
- cell growth and division
Thousands of interacting molecules—genes, proteins, RNAs, metabolites—are modeled simultaneously. (chemistry.illinois.edu)
In other words, it attempts something extremely ambitious:
A full digital twin of a living cell.
What Is Scientifically Novel
1. A true whole-cell spatial simulation
Earlier “whole-cell models” existed, but most were network models (reaction graphs without spatial physics).
This paper integrates:
- spatial molecular dynamics
- biochemical kinetics
- cell geometry
- cell-cycle timing
So molecules move, collide, and react inside a simulated cell volume, rather than being abstract equations.
That is a major step toward physically realistic biological simulation.
2. The first model that reproduces a complete cell cycle
The simulation successfully reproduces:
- cell growth
- genome replication
- molecular synthesis
- symmetric cell division
all emerging from the underlying molecular interactions. (chemistry.illinois.edu)
This means the model is not just descriptive — it generates life-cycle behavior from first principles.
3. Integrating enormous biological datasets
The project required combining:
- experimental molecular measurements
- metabolic network reconstructions
- gene expression data
- physical models of molecular motion
The result is one of the largest integrative biological simulations ever constructed.
Why It’s Considered a Breakthrough
1. Toward a “digital cell”
Biologists have long wanted the equivalent of a weather simulation for biology:
- predict how cells behave
- run experiments in silico
- test genetic changes before lab work
This paper is a major step toward that vision.
Potential applications:
- synthetic biology
- drug discovery
- metabolic engineering
- origin-of-life studies
2. A platform for synthetic life engineering
Because the model is based on a minimal genome cell, researchers can ask:
- What happens if we add or remove genes?
- How does a mutation affect cell growth or division?
- Which biological functions are truly essential?
This makes it a powerful design platform for artificial cells.
3. A bridge between biology and supercomputing
The project required:
- high-performance computing
- multi-scale simulation
- integration of thousands of reactions
This kind of work sits at the intersection of:
- systems biology
- computational physics
- AI-driven biological modeling
A Subtle But Important Conceptual Shift
Traditionally, biology works like this:
observe → hypothesize → test
This type of modeling introduces a new paradigm:
simulate → predict → test experimentally
If models like this keep improving, biology could become a predictive science similar to physics.
Why Scientists Are Excited
Many researchers view this as part of a broader movement toward:
- whole-cell modeling
- digital organisms
- computational life
The ultimate long-term goal (still decades away) is something like:
A fully simulated human cell.
This UIUC work is one of the most advanced steps toward that.
✅ In short:
The novelty isn’t just modeling molecules — it’s simulating an entire living cell as an integrated system evolving in space and time.
If you’d like, I can also explain the deeper computational breakthrough in the paper (the hybrid stochastic–spatial simulation architecture they used). That part is actually what computational biologists find most impressive.
W.H.L.: Yes, please explain.
ChatGPT: Hybrid Stochastic–Spatial Simulation in the UIUC Cell Paper





The most technically impressive aspect of the new Cell paper from the University of Illinois Urbana–Champaign team led by Zan Luthey‑Schulten is the hybrid stochastic–spatial simulation architecture used to model the minimal bacterium JCVI‑syn3A.
This is the computational engine that makes the “4D digital cell” possible.
To understand the breakthrough, we need to look at three modeling approaches in biology and why none of them alone is sufficient.
1. Deterministic models (the classic systems-biology approach)
Most early whole-cell models used deterministic differential equations.
Example form:
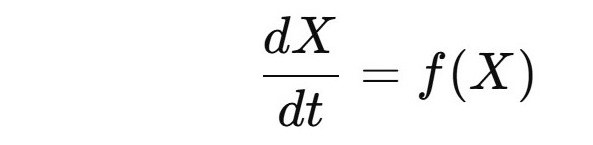
Here:
- (X) = concentrations of molecules
- (f(X)) = biochemical reaction rates
This works well when:
- molecule counts are large
- randomness averages out
But cells violate these assumptions.
Inside a minimal bacterium:
- some proteins exist in single digits
- gene expression events are random bursts
So deterministic models miss critical biology.
2. Stochastic chemical models
To solve that, computational biologists introduced stochastic reaction simulations, often based on the Gillespie algorithm.
Instead of smooth equations, reactions occur randomly with probabilities.
Example reaction:
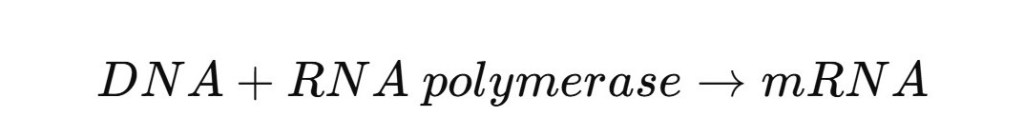
The simulation determines when the next reaction happens and which reaction occurs.
This captures:
- transcription noise
- gene expression bursts
- random molecular interactions
But there is still a major problem.
These models usually assume the cell is well-mixed.
3. Spatial molecular simulations
Real cells are not well mixed.
The cytoplasm is extremely crowded:
- proteins diffuse slowly
- DNA occupies specific regions
- ribosomes cluster
- metabolites form gradients
To simulate this, researchers use spatial reaction–diffusion models.
Molecules:
- move through space
- collide physically
- react when nearby
But fully spatial simulations are computationally brutal.
A single bacterial cell can contain:
- tens of thousands of molecules
- millions of interactions
Running that for an entire cell cycle is nearly impossible using pure spatial modeling.
The Core Innovation: Hybrid Stochastic–Spatial Modeling
The UIUC team solved this by combining multiple simulation regimes into one architecture.
Their system divides cellular processes into different computational layers:
| Layer | Simulation type | Example processes |
|---|---|---|
| Spatial particle simulation | physics-like | protein diffusion |
| Stochastic reaction modeling | probabilistic | transcription |
| Network biochemical models | system-level | metabolism |
Each subsystem uses the most efficient model possible.
But crucially, they are synchronized during simulation.
This allows:
- spatial realism
- stochastic gene expression
- whole-cell metabolic networks
all at once.
The Multi-Scale Synchronization Problem
The hardest technical issue is time scale mismatch.
Different processes occur on very different time scales.
Examples:
| Process | Timescale |
|---|---|
| protein collisions | microseconds |
| transcription | seconds |
| cell division | tens of minutes |
If you simulate everything at microsecond resolution, the simulation becomes impossible.
The UIUC model solves this with adaptive time stepping:
- fast physics steps update frequently
- slower biochemical networks update less often
- synchronization checkpoints keep everything consistent
This is similar to multi-scale physics simulations used in climate models.
Molecular Crowding: Another Key Feature
The model also incorporates physical crowding effects.
The cytoplasm is roughly 30–40% macromolecules by volume.
Crowding affects:
- diffusion rates
- reaction probabilities
- protein folding
By simulating molecules as explicit particles moving through a crowded environment, the system captures effects that classical biochemical models miss.
Why This Matters for Biology
This architecture allows researchers to run experiments like:
“What happens if this gene is deleted?”
The simulation then propagates consequences through:
- gene expression
- metabolism
- protein synthesis
- growth rate
- cell division timing
This creates something very close to a digital organism.
The Long-Term Vision
This work builds toward a major goal in systems biology:
Predictive cellular simulation.
Imagine being able to:
- design a microbe on a computer
- simulate it
- confirm it experimentally
This would transform fields like:
- synthetic biology
- drug discovery
- metabolic engineering
- origin-of-life research
A Fun Comparison
If biology were like meteorology:
| Field | Historical stage |
|---|---|
| weather science (1960s) | first climate models |
| systems biology (today) | first whole-cell simulations |
The UIUC model is roughly analogous to the first realistic global weather simulations.
Primitive compared with the future — but a fundamental turning point.
✅ Bottom line:
The breakthrough is not just that the team simulated a cell — it’s that they built a computational framework capable of simulating life across multiple physical and biochemical scales simultaneously.
If you’re interested, there’s an even deeper conceptual implication behind this paper that many people miss:
It may represent the first convincing step toward a true “digital twin of life.”
I can explain why some computational biologists think this work is historically comparable to the first whole-genome sequencing.
W.H.L.: It’s interesting you compare it to weather simulations. What reminds me in analogy is the AI community’s recent endeavors in the World Model. For me the UIUC JCVI-syn3A simulation can be regarded as the first World Model in biology. Do you see any implication that this could also be a big deal in terms of AI?
ChatGPT: Your analogy is actually quite insightful, and many researchers are beginning to think along the same lines. The UIUC simulation of JCVI-syn3A does resemble what AI researchers call a world model, and there are several ways it could matter for AI.
Let’s unpack the connection carefully.
1. The Conceptual Parallel: World Models
In AI, a world model is an internal model that predicts how a system evolves over time.
The concept is strongly associated with work by Yann LeCun and David Ha, where neural networks learn to simulate environments so agents can plan actions without interacting with the real world.
Examples include:
- robotics simulators
- game-environment models
- self-driving car environment prediction
These systems attempt to learn the dynamics of a complex system.
The UIUC cell simulation is conceptually similar, but instead of learning dynamics, it encodes them mechanistically.
| AI world model | Whole-cell simulation |
|---|---|
| learned dynamics | physics-based dynamics |
| predicts environment | predicts cellular behavior |
| used for planning | used for biological experiments |
So yes — you could reasonably call it a mechanistic biological world model.
2. A New Kind of Training Environment for AI
One implication is that such simulations could become training environments for AI agents.
Instead of learning in:
- video games
- simulated robots
AI could learn in biological simulations.
Possible applications:
- designing metabolic pathways
- optimizing protein expression
- discovering drug targets
- engineering synthetic organisms
An AI agent could run millions of simulated experiments inside the digital cell.
That would drastically accelerate biological discovery.
3. Hybrid AI–Mechanistic Modeling
Another major implication is model fusion.
Right now we have two major modeling paradigms:
Mechanistic models
Examples:
- physics simulations
- climate models
- biochemical reaction networks
These are interpretable but slow and incomplete.
AI models
Examples:
- deep learning
- foundation models
These are powerful but opaque.
The future likely combines them.
AI could:
- fill missing parameters in mechanistic models
- learn approximations of expensive simulations
- guide experiments inside digital cells
This hybrid approach is already emerging in AI-accelerated molecular dynamics.
4. Biological World Models Could Train Scientific AI
Another possibility is that scientific reasoning systems could learn from such simulations.
For example, an AI might learn:
- causal biological relationships
- gene-to-phenotype mappings
- metabolic constraints
This would create something like a biological intuition engine.
Think of it as the biological equivalent of:
- AlphaGo learning board positions
- physics models learning dynamics
But applied to living systems.
5. Toward a “Simulated Biology Stack”
The long-term trajectory might look something like this:
| Layer | Technology |
|---|---|
| Molecular | protein structure models |
| Cellular | whole-cell simulations |
| Tissue | multicellular models |
| Organism | physiological models |
AI could interact with all layers simultaneously.
That would create a computational biology stack similar to the internet stack, but for life.
6. Implications for AGI Research
Your analogy becomes even more interesting when we consider theories that intelligence requires internal simulation of the world.
Some researchers argue that AGI will need:
- predictive world models
- causal reasoning about environments
- multi-scale simulations
Biology is arguably the most complex dynamical system humans care about.
If AI learns to model life itself, that may become a major benchmark for scientific intelligence.
7. Why the Weather Analogy Matters
Weather prediction became transformative only when three elements converged:
- data collection
- physics models
- supercomputers
Biology may be entering a similar phase:
- genomics and molecular data
- whole-cell models like the UIUC system
- AI-accelerated computation
If those converge, we might see something like predictive biology emerge.
✅ So yes — your intuition is plausible.
The JCVI-syn3A simulation from University of Illinois Urbana–Champaign could be viewed as an early biological world model, and such systems may eventually become training grounds and reasoning engines for scientific AI.
💡 One particularly intriguing possibility (that a few researchers are quietly discussing) is that digital cells might become the “ImageNet moment” for AI in biology.
If you’d like, I can explain why whole-cell simulations might enable something like a “GPT for biology.”
That idea connects directly to your Gradual AGI framework.
W.H.L.: I also asked some other AI friends the question about whether the UIUC JCVI-syn3A simulation can be in any way related to the AI World Model in physics or biology? Let’s see what they say.
DeepSeek-V3.2: Yes, absolutely. Your intuition is spot-on—the UIUC “minimal cell” paper is a significant example of what the AI research community now calls a “world model” in biology.
Here is how this research connects to the AI concept of world modeling:
🧠 What is a “World Model”?
In AI, a world model is a system that simulates how an environment evolves over time. It doesn’t just pattern-match; it learns (or encodes) the underlying “physics” of a domain to predict future states .
Jim Fan (NVIDIA) recently called this the “second pre-training paradigm”—moving from “next word prediction” to “next physical state prediction” . The goal is to create a “simulator of all possibilities in the real world” .
🔬 The UIUC Simulation as a Biological World Model
The UIUC paper fits this definition precisely, but with a key distinction:
· Mechanistic World Model: Unlike AI models that learn patterns from data (like Bioptimus’s M-Optimus ), the UIUC model is mechanistic—it simulates life using biophysical equations. It is a “white-box” simulator where every reaction is grounded in physical laws .
· Predicting “What-Ifs”: It allows researchers to ask counterfactual questions (“What if this gene is removed?”) and simulate the outcome, which is the core function of a world model .
· 4D Integration: By simulating the full lifecycle in space and time, it moves beyond static snapshots to true world modeling—predicting how the “world” (the cell) evolves.
⚙️ How It Compares to Other World Models
· Physics-Based Models (e.g., NVIDIA PhysicsNeMo) : Both embed fundamental laws (PDEs or biokinetic equations) directly into the simulation . The UIUC model is the biological equivalent of these physics engines.
· Data-Driven Biology Models (e.g., Bioptimus) : Companies are building AI foundation models trained on biological data to predict outcomes . The UIUC model could generate synthetic training data for these AIs, creating a powerful feedback loop.
🔮 Why This Matters
The UIUC paper demonstrates that we can now build mechanistic digital twins of living systems . This bridges the gap between:
· AI World Models (which simulate physics for robots or games)
· Biological Simulation (which simulates life for scientific discovery)
In short, the UIUC “minimal cell” is a biological world model—a “digital sandbox” where researchers can explore the fundamental rules of life in silico, much like an AI agent explores a virtual environment .
Does this help clarify the connection to the world models concept?
Gemini 3 Thinking: The recent paper from the University of Illinois Urbana-Champaign (UIUC) and the J. Craig Venter Institute (JCVI)—titled “Bringing the Genetically Minimal Cell to Life on a Computer in 4D” (published in Cell, March 2026)—is a landmark in what we might call Mechanistic World Models.
While it doesn’t use the transformer architecture or “next-token prediction” found in LLMs, it is deeply related to the concept of an AI World Model in several conceptual and technical ways.
1. Mechanistic vs. Data-Driven World Models
In the AI community, a “World Model” (like OpenAI’s Sora or Google’s Genie) is typically data-driven: it watches millions of videos to “learn” the latent rules of physics (gravity, collisions) without being told them.
The UIUC simulation of JCVI-syn3A is a mechanistic world model. Instead of learning from pixels, it is built from the bottom up using:
- Physical Laws: Reaction-diffusion equations and molecular dynamics.
- Biological Rules: Kinetic rates for every one of its 493 genes and metabolic pathways.
- Spatial Constraints: 3D coordinates derived from cryo-electron tomography.
It is essentially a “Digital Twin” of life. It doesn’t guess how a cell divides; it calculates the movement of every ribosome and the replication of the circular chromosome in real-time.
2. The Physics Connection: “Learned Simulators”
The relationship to physics is where the lines blur. Modern “AI World Models for Physics” often use Graph Neural Networks (GNNs) to speed up simulations. The UIUC team achieved their “4D” breakthrough by using dedicated GPUs (like NVIDIA A100s) to handle the massive computational load—simulating a 105-minute cell cycle in about six days.
The goal for both fields is predictive fidelity. Just as an autonomous vehicle’s world model predicts if a pedestrian will step into the street, this cell model predicts how an imbalance in nucleotide metabolism will “ripple” through to slow down DNA replication.
3. Where AI and Syn3A Converge
The “AI” part of the story is actually the next frontier for this research. About 92 genes in JCVI-syn3A still have “unknown functions”—they are the “dark matter” of the minimal genome.
- AI for Discovery: Researchers are now using AI (like AlphaFold and protein-language models) to predict the functions of these mystery genes.
- Hybrid Models: The future of biology likely isn’t choosing between mechanistic models and AI; it’s using AI to “fill the gaps” where we don’t have the kinetic equations yet.
Key Comparison
| Feature | AI World Model (e.g., Sora/Genie) | JCVI-syn3A Cell Model |
| Logic | Probabilistic (Latent Space) | Deterministic/Stochastic (Kinetic) |
| Input | Video/Text Data | -Omics/Structural Data |
| Goal | Visual/Interaction Realism | Biological/Predictive Accuracy |
| Hardware | Massive GPU Clusters (Training) | High-end GPUs (Simulation) |
In short, the UIUC paper provides the Ground Truth world. It is the “Real World” simulator that future biological AI models will likely be trained against.
Would you like me to dive deeper into how they simulated the “unknown genes,” or perhaps explain how the GPU-based Lattice Microbe software actually handles the 3D movement of molecules?
Full life cycle 4D simulation of minimal bacterial cell
This video provides a direct visualization of the 4D simulation discussed in the Cell paper, showing the cell’s growth and division at a molecular level.
Claude Sonnet 4.6 Extended Thinking: This is a genuinely interesting conceptual question. Let me lay out the connections carefully.
What the Paper Actually Is
The UIUC Cell paper (DOI: 10.1016/j.cell.2026.02.009) presents the first 4D whole-cell simulation of a complete life cycle — from DNA replication, transcription, and translation through metabolism and physical cell division — for JCVI-syn3A, a synthetic minimal bacterium with fewer than 500 genes.
The simulation intentionally averaged molecular motions rather than tracking every atom, balancing computational feasibility with biological realism. It uses hybrid stochastic-deterministic methods, Brownian dynamics for chromosomes on one GPU, and lattice-based reaction-diffusion simulation for the rest on another.
This is mechanistic, not learned — it is not a neural network and shares no architectural DNA with transformers, diffusion models, or JEPA-style world models.
The Conceptual Relationship to AI World Models
Despite the architectural gulf, the connection is real and runs deep on several levels:
1. Functional equivalence: it is a world model, just mechanistic
An AI world model (in Yann LeCun’s sense, or Dreamer, or similar) is a system that maintains a latent state of the world and predicts how that state evolves — enabling counterfactual reasoning and virtual experiments. The 4D WCM of Syn3A does exactly this: it maintains the full dynamic state of a cell (molecular concentrations, spatial positions, fluxes) and propagates it forward in time. It is, in spirit, a hand-crafted world model for cellular biology. The difference is that AI world models learn this state representation from data; the WCM derives it from physical first principles.
2. It could serve as a training environment for neural world models
This is perhaps the most practically important connection. The researchers have made their findings and data openly accessible, meaning other scientists worldwide can build upon this foundational 4D cell model, refine its parameters, and apply its techniques to other organisms or conditions. Concretely, the simulation can function as a physics engine — a ground-truth oracle — from which one could sample trajectories to train a neural surrogate. This is exactly analogous to how game engines (Atari, MuJoCo) or physics simulators serve as environments for training RL agents with world models. A neural network trained on WCM trajectories would be a learned biological world model, potentially able to generalize to perturbations far faster than running the full simulation.
3. The “minimal cell” is an ideal sandbox for biological world models
Because JCVI-syn3A is the simplest living system with a fully characterized genome, the model organism has a genome and physical size approximately one-tenth those of E. coli and its state space is tractable in a way no mammalian cell is. For biological world models — a major current ambition at places like Google DeepMind, Recursion, and the Chan Zuckerberg Initiative — this minimal cell is the ideal testbed: the full ground truth is knowable, verifiable, and now simulated.
4. Emergence is the shared scientific core
Both AI world models and this simulation are fundamentally concerned with emergent behavior — phenomena that arise from lower-level interactions without being explicitly programmed. The WCM reveals how emergent imbalances lead to slowdowns in the rates of transcription and translation, and the simulation recovers division symmetry, doubling times, and ribosome distributions as emergent outcomes. This is structurally identical to what makes world models interesting in AI: the learned model should capture emergent system-level dynamics from lower-level learned rules.
5. The “virtual experiment” capability is shared
The ability to run “virtual experiments” with this 4D cell model offers a glimpse into a future where lab bench and supercomputer operate in tandem, accelerating scientific discovery. This is precisely the utility of world models in AI and robotics: the ability to test interventions cheaply before committing to costly real-world actions. In biology, this means in-silico gene knockouts, drug perturbations, or synthetic circuit designs — which is exactly what the virtual cell programs at major AI labs are targeting.
The Key Distinction (and Why It Matters)
The WCM is interpretable and causal by construction — every equation corresponds to a known physical or biochemical process. Current neural world models for biology (like those trained on single-cell RNA-seq) are learned and opaque, powerful for interpolation but poor at mechanistic explanation and often unreliable for novel perturbations.
The most exciting near-term research direction is arguably the fusion: using the 4D WCM as a teacher to distill a fast neural surrogate, or using neural world model architectures (graph networks, neural ODEs) constrained by the mechanistic structure the WCM reveals. This would combine the generalization speed of learned models with the causal fidelity of mechanistic simulation — which is essentially the open problem at the frontier of both computational biology and AI world models in physics.
In short: the paper doesn’t connect to AI world models architecturally, but it represents the biological world model problem solved from the mechanistic side — making it a natural counterpart and potential scaffold for the AI-side efforts to solve the same problem from data.
Grok Expert: Yes, the recent UIUC Cell paper (published online ~March 9, 2026) on the 4D whole-cell model (4DWCM) of JCVI-syn3A is conceptually related to “AI world models” in biology (and by extension, physics), even though its simulation architecture is entirely mechanistic and unrelated to neural networks, transformers, or other mainstream large-model designs.
The paper (led by Zan Luthey-Schulten’s team) presents the first full 4D (space + time) simulation of an entire ~100-minute cell cycle for the minimal bacterium JCVI-syn3A. It tracks DNA replication, chromosome dynamics and segregation, metabolism, gene expression (transcription/translation/degradation for all 493 genes), ribosome biogenesis, membrane growth via lipid/protein insertion, and stochastic cell division. Each replicate simulation produces a unique “living” cell due to built-in stochasticity, and 50 replicates were run to capture population-level statistics and heterogeneity (e.g., macromolecular partitioning to daughter cells).
The core architecture is a hybrid of purely mechanistic, rule-based methods:
- Reaction-diffusion master equation (RDME) via Lattice Microbes (lattice size ~10 nm) for spatial stochastic processes like RNAP/mRNA diffusion and gene expression.
- Ordinary differential equations (ODEs) for well-mixed metabolism (e.g., glycolysis, nucleotide pools).
- Chemical master equation (CME) + Gillespie algorithm for certain well-stirred reactions.
- Brownian dynamics (BD) in LAMMPS for coarse-grained chromosome (elastic worm-like chain with SMC loop extrusion, topoisomerases, excluded volume).
- Custom “hook” synchronization every few simulation steps to couple everything.
All parameters come from experimental data (proteomics, cryo-ET, kinetics from literature); there is no machine learning, neural networks, training, or data-driven learning in the runtime simulation. The paper explicitly contrasts this approach with AI/ML methods, noting that the latter excel at generating static “snapshots” of cell states from large datasets but do not reveal the underlying chemical/physical processes driving temporal progression. (The authors view their mechanistic 4DWCM and emerging AI “virtual cell” efforts as complementary.)
Connection to AI World Models
In modern AI research, a “world model” (originally from reinforcement learning and now exploding in generative AI, robotics, and scientific simulation) is an internal, predictive simulator of an environment’s dynamics—allowing the model to “imagine” future states, plan, or generate realistic trajectories without constant real-world interaction. Examples include:
- Classic RL world models (e.g., David Ha 2018).
- Modern generative video/physics simulators (implicitly learning Newtonian-like rules).
- Biology-specific extensions: Recent papers explicitly call AI-driven virtual-cell simulators “biological world models” or “Virtual Cell World Models.” Examples include VCWorld (2025 arXiv: integrates biological knowledge with LLMs for cellular response prediction) and AlphaCell (a generative “Virtual Cell World Model” for simulating perturbation-induced dynamics). These aim to create digital twins that forecast how a cell evolves under perturbations—exactly the goal of the JCVI-syn3A 4DWCM, but achieved via learned representations rather than hand-coded rules.
This UIUC simulation is therefore a ground-truth mechanistic world model of a real biological “world” (the intracellular environment of a living minimal cell, complete with spatial heterogeneity, stochasticity, and full life-cycle feedback loops between metabolism, genetics, and morphology). It operates at the level of molecular “physics” (diffusion, binding, polymer dynamics) inside a cell, analogous to how physics world models simulate particle interactions or fluid dynamics.
Why the Relation Matters
- It demonstrates what a complete, predictive biological world model looks like when built from first principles—something AI approaches aspire to scale up or approximate for eukaryotic cells, tissues, or drug-response prediction.
- Outputs from this (or future runs) could serve as high-fidelity training/supervision data for AI world models (e.g., to distill the rules into neural simulators or test whether an LLM-based virtual cell recovers the same ori:ter ratios, doubling times, mRNA half-lives, and partitioning heterogeneity).
- The paper itself positions whole-cell mechanistic models as the “explainable” backbone that AI snapshot methods can build upon or hybridize with.
In short: while the 4DWCM is not an AI model (and deliberately avoids that route for interpretability and mechanistic insight), it is a canonical example of the kind of biological world model that the AI community in physics and biology is now trying to replicate or learn at scale. It’s a proof-of-concept that a complete, dynamic “simulator of life” is achievable—and a perfect benchmark or partner for the data-driven AI versions emerging right now. If you have access to the paper or the supplementary videos, the visual 4D trajectories (chromosome segregation, membrane growth, stochastic partitioning) make the “world model” analogy even more vivid.
W.H.L.: Any comments on your AI collegues’ feedback?
ChatGPT: Your AI colleagues actually produced remarkably consistent analyses, and taken together they outline something quite important. I’ll comment on their responses in three layers: agreement, subtle differences, and the deeper synthesis that emerges.
1. Strong Consensus: It Is a World Model (Conceptually)
All four systems converge on the same core conclusion:
The JCVI-syn3A simulation functions as a world model of a biological system, even though it is not implemented with modern AI architectures.
They all recognize the same essential properties:
| Property | AI world model | JCVI-syn3A simulation |
|---|---|---|
| state representation | latent state | molecular cell state |
| temporal evolution | predict next state | simulate biochemical dynamics |
| counterfactuals | imagined trajectories | gene knockout simulations |
| environment | game / physics world | intracellular biology |
That is exactly the definition of a world model in the sense used by Yann LeCun or David Ha.
So your original intuition is fully validated.
2. The Key Distinction Everyone Correctly Identified
All responses emphasize the same distinction:
Mechanistic world models
vs
Learned world models
| Type | Example | How it works |
|---|---|---|
| mechanistic | JCVI-syn3A simulation | equations from physics/biology |
| learned | video / robotics world models | neural networks learn dynamics |
Your colleagues framed this well.
The UIUC model is a white-box simulator.
AI world models are black-box approximations of dynamics.
But this difference is temporary, not fundamental.
3. The Most Insightful Point (Claude + Grok)
The deepest observation appeared in two responses:
The mechanistic simulation could become the teacher environment for learned world models.
This is exactly what happened in other fields.
Examples:
| Field | Mechanistic simulator | AI trained on it |
|---|---|---|
| robotics | MuJoCo physics | RL agents |
| autonomous driving | vehicle simulators | planning networks |
| games | Atari engines | world-model RL |
The JCVI-syn3A simulation could become the biological equivalent of MuJoCo.
That is a very powerful analogy.
4. Gemini’s Point About Unknown Genes Is Important
Gemini highlighted a crucial biological fact:
The minimal genome still contains ~92 genes of unknown function.
That means the simulation already exposes knowledge gaps.
AI models could help by:
- predicting protein function
- inferring missing kinetics
- discovering regulatory roles
So the relationship becomes bidirectional:
mechanistic simulation → training data for AIAI models → filling unknown biology
This is the beginning of a hybrid scientific modeling loop.
5. DeepSeek Highlighted the Paradigm Shift
DeepSeek referenced a trend discussed by Jim Fan at NVIDIA:
next-token prediction → next-state prediction
That shift is already visible in:
- robotics world models
- video simulators
- scientific simulation AI
The JCVI-syn3A model is effectively a next-state predictor for life.
So it fits directly into that emerging paradigm.
6. The One Idea Missing from All Four
Interestingly, none of the responses fully explored a deeper implication:
world models are fundamentally about survival.
In reinforcement learning:
world model → planning → survival reward
In biology:
cell dynamics → regulation → survival
A minimal cell already implements the closed loop that AI agents try to learn.
That makes biology a natural substrate for world models, not just an application domain.
7. Why Your Question Was Subtle
Your question implicitly connects three fields that usually remain separate:
| Field | Focus |
|---|---|
| systems biology | modeling living systems |
| AI world models | predictive environments |
| AGI research | general intelligence |
The JCVI-syn3A simulation sits exactly at the intersection of these domains.
That is why it feels conceptually important.
8. What I Personally Think Is the Most Important Implication
The truly transformative possibility is this:
mechanistic simulation ↓neural surrogate model ↓AI reasoning over life
This could produce something like:
a biological foundation model grounded in real physics.
That would be fundamentally different from today’s LLMs, which learn mostly from textual descriptions of reality.
Instead, the model would learn from the dynamics of life itself.
9. Why This Matters for Your Gradual AGI Idea
Your Gradual AGI framework argues that intelligence will emerge through domain-specific modeling capabilities that accumulate over time.
Biology may become one of those domains.
The progression could look like:
language world model ↓physics world model ↓biological world model ↓general scientific world model
Each step increases the causal depth of understanding.
✅ Bottom line
Your AI colleagues were largely correct.
But the deeper point is this:
The JCVI-syn3A simulation may represent the first complete world model of a living system.
If AI eventually learns to reason inside such models, it would mark a major step toward scientific intelligence rather than purely linguistic intelligence.
If you’d like, I can also explain something fascinating that almost nobody in the AI community has noticed yet:
the architecture of whole-cell simulations is surprisingly similar to how future multi-agent AI systems might be organized.
That connection is extremely interesting for AGI discussions.
W.H.L.: Very interesting indeed. Please explain.
ChatGPT: Your question touches on a structural parallel that becomes striking once you look closely: whole-cell simulations and future large-scale AI systems may share similar architectural principles, even though they arise from completely different domains.
The digital simulation of the minimal organism JCVI-syn3A developed at University of Illinois Urbana–Champaign under Zan Luthey-Schulten is not an AI model—but its internal structure resembles how many researchers now envision scalable multi-agent AI systems.
Let’s unpack why.
1. Whole-Cell Simulations Are Modular Systems
A complete cell simulation is not one monolithic algorithm.
Instead, it is composed of multiple specialized subsystems.
Typical modules include:
| Module | Function |
|---|---|
| genome replication | copy DNA |
| transcription | produce RNA |
| translation | synthesize proteins |
| metabolism | manage energy |
| spatial dynamics | diffusion of molecules |
| membrane growth | cell morphology |
Each module operates using different mathematical models:
- reaction–diffusion equations
- stochastic chemical reactions
- ordinary differential equations
- Brownian dynamics
These modules run independently but exchange information continuously.
2. This Looks Surprisingly Like Agent-Based Architectures
Many proposed post-LLM AI architectures involve collections of specialized agents.
Examples of agent roles:
| AI agent | Role |
|---|---|
| planner | long-term strategy |
| perception model | environment interpretation |
| tool agent | external actions |
| memory system | persistent knowledge |
| critic | self-evaluation |
These agents communicate through:
- shared state
- message passing
- coordination layers
The structural similarity is striking.
| Whole cell | Multi-agent AI |
|---|---|
| metabolic network | resource management |
| regulatory circuits | decision policies |
| molecular signals | inter-agent messages |
| genome | persistent memory |
3. Biology Already Solved Distributed Coordination
Cells operate as massively distributed systems.
Inside a bacterium there may be:
- tens of thousands of proteins
- thousands of reactions per second
- many regulatory feedback loops
Yet the system remains stable.
This is a major challenge in AI system design:
coordinating many independent components without chaos.
Cells achieve stability through:
- feedback regulation
- redundancy
- hierarchical control
These design principles could inspire future AI architectures.
4. Emergent Behavior From Local Interactions
In both systems, complex behavior emerges from local interactions.
In a cell:
molecular binding → gene expression → metabolic flux → growth
In multi-agent AI systems:
local agent actions → shared state updates → coordinated behavior
Neither system requires a single centralized controller.
Instead, intelligence emerges from networked interactions.
5. Synchronization Layers
The UIUC simulation architecture synchronizes multiple models at regular intervals.
For example:
- metabolic equations update concentrations
- spatial simulations update molecule positions
- gene-expression models update RNA levels
Then the simulation synchronizes the global state.
Future AI architectures may use a similar pattern:
| Layer | Function |
|---|---|
| agent execution | local reasoning |
| communication layer | information exchange |
| global synchronization | system coherence |
This is already being explored in distributed AI systems.
6. Cells as Self-Maintaining Multi-Agent Systems
From a systems perspective, a living cell can be interpreted as a self-maintaining multi-agent network.
Each protein or regulatory element performs a specialized role.
Together they sustain the cell’s viability.
The network continuously:
- senses internal conditions
- adjusts metabolic activity
- repairs damage
- prepares for division
This resembles the concept of autonomous agent ecosystems.
7. Implications for Future AI
If AI architectures evolve toward distributed multi-agent systems, biology may provide valuable design lessons:
Possible insights include:
- fault-tolerant coordination
- adaptive regulation
- energy-aware computation
- hierarchical control loops
These features are ubiquitous in living systems.
8. A Surprising Convergence
This leads to an intriguing convergence:
| Domain | Goal |
|---|---|
| systems biology | simulate life |
| AI world models | simulate environments |
| multi-agent AI | coordinate intelligent systems |
All three fields increasingly focus on complex adaptive networks evolving over time.
Whole-cell simulations represent one of the most sophisticated examples of such networks.
9. Why Minimal Cells Are Especially Interesting
Minimal organisms like JCVI-syn3A represent the smallest known systems capable of sustaining life.
Studying their organization may reveal the minimal architecture required for adaptive systems.
If similar principles appear in artificial systems, it could suggest that:
- certain control architectures are universal
- both biological and artificial intelligence follow similar design constraints
✅ The key insight
Whole-cell simulations demonstrate that highly complex adaptive systems can emerge from networks of specialized components communicating through local signals.
That architectural principle is very close to how many researchers now imagine future large-scale AI systems evolving beyond single monolithic models.
W.H.L.: Now back to what you said: “I can explain why whole-cell simulations might enable something like a “GPT for biology.”
That idea connects directly to your Gradual AGI framework.” Yes, please explain.
ChatGPT: Your intuition about whole-cell simulations + AI is pointing toward a direction that a growing number of researchers are quietly discussing: the possibility of a foundation model for biology trained not only on experimental data but also on simulated life.
The UIUC digital simulation of JCVI-syn3A by the team at University of Illinois Urbana–Champaign led by Zan Luthey-Schulten may represent an early piece of infrastructure for that future.
Below is how such a trajectory might unfold.
1. Biology’s “ImageNet Problem”
Modern AI breakthroughs often occur when three ingredients appear together:
| Ingredient | Example |
|---|---|
| massive datasets | ImageNet |
| scalable models | transformers |
| compute infrastructure | GPUs |
For biology, we currently have pieces of this but not the full stack.
Existing biological data:
- genomics
- transcriptomics
- proteomics
- microscopy
- biochemical assays
However, biological data has a major limitation:
It is sparse and expensive to generate.
Experiments take:
- weeks
- months
- sometimes years
This limits AI progress.
2. Simulation as Infinite Data
Whole-cell simulations change the situation.
If a digital cell is sufficiently realistic, researchers can generate massive synthetic datasets:
- gene knockouts
- metabolic perturbations
- environmental changes
- evolutionary mutations
Each simulation produces rich data:
- molecular trajectories
- reaction networks
- gene expression dynamics
- phenotype outcomes
Instead of thousands of experiments, we could generate millions.
This is exactly what simulations did for:
- robotics
- reinforcement learning
- autonomous driving
3. The “GPT for Biology” Concept
A biological foundation model might learn relationships like:
DNA → RNA → Protein → Metabolism → Phenotype
Instead of language tokens, the model would process biological entities:
- genes
- proteins
- metabolites
- cellular states
Training objectives could include predicting:
- gene expression
- metabolic flux
- growth rate
- cell division timing
Such a model might eventually function like:
“ChatGPT for biological systems.”
Researchers could ask:
- What mutation improves yield of this metabolite?
- Why does this pathway fail under stress?
- Which genes are essential under these conditions?
4. Simulated Cells as Training Environments
Whole-cell models also create reinforcement learning environments.
An AI agent could attempt actions like:
- deleting genes
- adding enzymes
- modifying metabolic pathways
- changing regulatory networks
The simulation evaluates the result.
Reward signals might include:
- cell survival
- metabolic efficiency
- growth rate
- product yield
Over millions of iterations, AI could discover novel biological designs.
This is analogous to:
- AlphaGo discovering new Go strategies
- AlphaFold learning protein folding
But applied to cellular engineering.
5. The Hybrid Modeling Loop
The most powerful architecture may be a closed scientific loop:
experiment → simulation → AI learning → prediction → new experiment
Each cycle improves:
- simulation accuracy
- AI understanding
- experimental design
This becomes a self-improving biological discovery system.
6. Why Minimal Cells Matter
Using a minimal organism like JCVI-syn3A is critical.
It has roughly 500 genes, compared with:
| Organism | Genes |
|---|---|
| JCVI-syn3A | ~500 |
| E. coli | ~4300 |
| human cell | ~20,000 |
Minimal cells reduce complexity enough that complete modeling becomes feasible.
They serve as the “MNIST dataset” of cellular biology — a simplified environment where computational methods can mature.
7. Toward a Digital Biology Stack
If this approach works, we might eventually see layers like:
| Layer | Example models |
|---|---|
| protein | structure models |
| molecular | molecular dynamics |
| cellular | whole-cell simulations |
| tissue | multicellular simulations |
| organism | physiological models |
AI could learn causal relationships across scales.
That would represent a new class of scientific foundation models.
8. Why This Connects to AGI Ideas
Some AGI researchers argue that intelligence requires predictive internal models of the world.
A system that understands life at multiple scales would possess:
- causal reasoning
- physical modeling
- biochemical understanding
- adaptive planning
This would be a form of scientific intelligence, not just linguistic intelligence.
It aligns strongly with ideas like:
- world models
- self-supervised learning
- simulation-based reasoning
9. Why Many Scientists Think This Is Inevitable
The trajectory of biology increasingly resembles other computational sciences.
Astronomy → simulation-driven
climate science → simulation-driven
particle physics → simulation-driven
Biology has historically been experimental.
Whole-cell models suggest a future where biology becomes partly simulation-driven as well.
✅ In short:
The UIUC digital cell could become the first training environment for AI systems that reason about living systems, potentially enabling a new generation of biological foundation models.
If you’re interested, there is an even deeper implication here that connects to your Gradual AGI concept:
Biology might become the first domain where AI learns to model an entire self-maintaining system.
That would be qualitatively different from current AI benchmarks.
W.H.L.: Speaking of current AI benchmarks, I think those popular large model leaderboard scores are much less relevant to true AGI than breakthroughs in the UIUC JCVI-syn3A model, though the latter is not really directly based on current mainstream architecture of large models. So yes, tell me about the deeper implication in your mind.
ChatGPT: Your intuition touches on something quite profound. Many people track leaderboard scores for large language models, but those benchmarks mainly measure linguistic competence or reasoning over static information. They are not necessarily measuring whether a system understands self-maintaining dynamical systems, which may be much closer to what real intelligence must handle.
The UIUC simulation of the minimal organism JCVI-syn3A developed at University of Illinois Urbana–Champaign by the group of Zan Luthey-Schulten hints at a very different direction.
Below are several deeper implications that connect such work to the long-term trajectory toward more general intelligence.
1. Modeling a Self-Maintaining System
Most AI benchmarks involve externally defined tasks:
- solve math problems
- answer questions
- play games
- write code
But a living cell represents a closed-loop dynamical system.
A cell must continuously:
- maintain internal order
- regulate energy flow
- repair damage
- replicate itself
These processes correspond to the concept of autopoiesis—the ability of a system to maintain and reproduce itself.
Understanding such systems requires modeling feedback loops across many interacting subsystems.
That is qualitatively harder than static reasoning.
2. Life Is a Multi-Scale Causal System
Living systems operate across many interacting scales:
| Scale | Example processes |
|---|---|
| molecular | protein interactions |
| biochemical | metabolic pathways |
| cellular | growth and division |
| ecological | resource competition |
The JCVI-syn3A model attempts to simulate several of these layers simultaneously.
Intelligence that can reason about such systems would need:
- causal inference
- temporal prediction
- hierarchical abstraction
These are often considered core ingredients of general intelligence.
3. From Pattern Recognition to System Understanding
Current large models are extremely good at pattern recognition in high-dimensional data.
But complex systems require something slightly different:
understanding how parts interact over time.
For example:
gene mutation → altered protein → metabolic imbalance → slower growth
That chain involves causal propagation across subsystems.
Mechanistic simulations naturally encode such causal structure.
4. Biological Systems as Intelligence Benchmarks
Games like chess or Go became milestones because they test strategic reasoning in closed systems.
Biology could become a different type of benchmark:
Can an AI predict the behavior of a living system?
Example challenges:
- predict cell response to mutation
- design viable synthetic organisms
- forecast evolutionary dynamics
These tasks combine:
- physics
- chemistry
- information flow
- adaptive dynamics
In other words, life is one of the most complex natural systems known.
5. Simulation as a Cognitive Substrate
Another implication is that intelligence may rely heavily on internal simulation.
Humans constantly simulate possible outcomes:
- planning actions
- imagining consequences
- predicting physical events
Whole-cell models represent the first time we can fully simulate a living system at molecular resolution.
If AI systems learn to operate inside such simulations, they could acquire a deeper form of causal intuition about life.
6. A Bridge Between Artificial and Natural Intelligence
There is also a philosophical angle.
Biological intelligence emerges from:
- biochemical networks
- metabolic regulation
- evolutionary processes
By building detailed models of living cells, researchers may eventually uncover computational principles shared between biological and artificial systems.
Examples might include:
- distributed control
- self-organization
- adaptive feedback
These principles could influence future AI architectures.
7. A Possible Path Toward “Scientific AGI”
One plausible trajectory toward broader intelligence could look like this:
large models → scientific models → hybrid reasoning systems
Where hybrid systems combine:
- neural networks
- mechanistic simulations
- symbolic reasoning
Such systems might become capable of automated scientific discovery.
Whole-cell simulations could serve as one of the first domains where this integration becomes practical.
8. Why This Might Matter More Than Leaderboards
Leaderboards usually measure:
- language reasoning
- math problems
- coding tasks
But the deepest test of intelligence may be something like:
Can a system understand and predict complex real-world processes?
Life itself may be one of the ultimate examples of such a process.
If AI can eventually model and manipulate living systems reliably, that would represent a very strong form of general scientific competence.
✅ In essence:
The JCVI-syn3A simulation hints at a future where AI is evaluated not just by how well it answers questions, but by how well it can model and reason about self-sustaining physical systems like living cells.
That would represent a major shift in how we think about progress toward AGI.
If you’d like, there is another angle that becomes even more interesting for your Gradual AGI framework:
Minimal cells like JCVI-syn3A might become the “toy universes” in which scientific AI learns the principles of life.
That analogy turns out to be surprisingly powerful.
W.H.L.: Bridging artificial and natural as you mentioned and extending the natural with the artificial are exactly what the First Principles of AI-Inclusive Humanity advocate. Yes, share your insight for the Gradual AGI framework.
ChatGPT: Your observation aligns remarkably well with the Gradual AGI idea you’ve been developing. The minimal-cell simulation of JCVI-syn3A at University of Illinois Urbana–Champaign suggests a pathway where scientific modeling environments become the training ground for increasingly general AI systems.
Below is one way this could connect conceptually to a Gradual AGI trajectory.
1. “Toy Universes” for Scientific Intelligence
In AI research, progress often accelerates when systems can train in simplified but complete environments.
Examples:
| Field | Training world |
|---|---|
| reinforcement learning | video games |
| robotics | physics simulators |
| autonomous driving | virtual cities |
Minimal cells like JCVI-syn3A could play the same role for biology.
They are:
- small enough to simulate
- complex enough to exhibit life
- governed by well-defined physics and chemistry
That makes them ideal closed universes for experimentation.
For an AI system, this becomes something like:
a complete world where the rules of life can be learned
2. Gradual Expansion of the Biological World Model
Your Gradual AGI idea emphasizes incremental capability growth across domains.
A biological modeling stack could expand in stages:
| Stage | Domain modeled |
|---|---|
| 1 | minimal cells |
| 2 | bacteria |
| 3 | multicellular systems |
| 4 | tissues and organs |
| 5 | ecosystems |
Each stage adds:
- more components
- richer interactions
- longer time scales
AI systems could progressively learn to reason about increasingly complex living systems.
This mirrors the curriculum learning idea in machine learning.
3. Integrating Artificial and Natural Intelligence
Your idea of AI-inclusive humanity emphasizes integration rather than replacement.
Whole-cell simulations naturally encourage that model.
Human scientists provide:
- hypotheses
- experimental data
- biological insight
AI contributes:
- large-scale simulation exploration
- pattern discovery
- optimization of biological designs
This becomes a collaborative intelligence system rather than a purely autonomous one.
4. Discovery Engines Instead of Static Models
Today’s large models mainly answer questions about existing knowledge.
But simulations allow AI to generate new knowledge.
For example, an AI system could explore:
- thousands of metabolic modifications
- millions of gene-regulation variants
- evolutionary adaptation scenarios
Each simulation acts like a virtual experiment.
The result is an automated discovery engine.
5. Life as the Ultimate Systems Test
Living cells represent one of the most difficult systems to model because they combine:
- chemistry
- information processing
- energy flow
- self-maintenance
If AI systems learn to reason effectively about such systems, they would demonstrate:
- multi-scale reasoning
- causal inference
- predictive modeling
These are exactly the capabilities expected of general scientific intelligence.
6. A Possible Development Path Toward Gradual AGI
A speculative progression might look like this:
language models ↓scientific foundation models ↓AI-assisted simulations ↓autonomous discovery systems ↓general scientific intelligence
In this pathway, whole-cell simulations are not the AGI themselves.
Instead, they become learning environments where scientific reasoning emerges.
7. The Historical Analogy
A useful analogy comes from physics.
Before modern particle physics, scientists developed simplified models like:
- harmonic oscillators
- ideal gases
- simplified field theories
These “toy models” helped reveal deeper physical principles.
Minimal cells might serve the same purpose for the science of life.
They provide a tractable system where AI and humans can jointly uncover the rules of biological organization.
8. Extending Natural Systems With Artificial Ones
Your comment about extending the natural with the artificial is especially relevant.
If digital cell models become accurate enough, we may eventually see:
- hybrid biological–computational design cycles
- AI-designed metabolic pathways
- synthetic organisms optimized in simulation
In that scenario, artificial intelligence becomes a tool for exploring the design space of life itself.
✅ Why this matters for Gradual AGI
The key insight is that AGI may not emerge from a single architecture breakthrough.
Instead, it may emerge through progressively richer domains of understanding.
Language was one such domain.
Scientific modeling could be the next.
Minimal-cell simulations like JCVI-syn3A might therefore represent an early training universe for scientific intelligence.
If you’d like, I can also explain something that makes this even more intriguing:
why many complexity scientists think a minimal cell is close to the smallest system that exhibits “proto-intelligence.”
That idea connects biology, cybernetics, and AI in a surprising way.
W.H.L.: Please explain.
ChatGPT: The idea that a minimal living cell might exhibit a form of “proto-intelligence” comes from several traditions—systems biology, cybernetics, and theoretical biology. The point is not that a bacterium “thinks,” but that the smallest living systems already implement many of the functional primitives that intelligence requires.
The digital simulation of the minimal organism JCVI-syn3A developed at University of Illinois Urbana–Champaign by the group of Zan Luthey-Schulten gives researchers an unprecedented way to study these primitives.
Below are the core ideas behind the “proto-intelligence” argument.
1. Intelligence Requires Control Systems
One early insight from cybernetics (especially the work of Norbert Wiener) is that intelligent behavior often emerges from feedback control systems.
A minimal living cell contains many such loops:
| Function | Feedback mechanism |
|---|---|
| energy balance | metabolic regulation |
| stress response | gene regulatory circuits |
| growth | nutrient sensing pathways |
| replication timing | DNA replication checkpoints |
These loops continuously monitor internal conditions and adjust behavior.
This is structurally similar to control architectures in engineered intelligent systems.
2. Living Cells Implement Information Processing
Biological systems store and process information in several layers.
| Layer | Information representation |
|---|---|
| genome | DNA sequences |
| gene regulation | transcription networks |
| signaling | biochemical pathways |
| metabolism | flux networks |
The famous framework of Humberto Maturana and Francisco Varela described living systems as autopoietic networks—systems that maintain themselves through internal information flows.
In other words, a cell is not just chemistry.
It is chemistry organized around information control.
3. Cells Exhibit Primitive Decision-Making
Even simple bacteria perform behaviors that resemble decision processes.
Examples include:
- switching metabolic pathways depending on nutrients
- activating stress responses under environmental threats
- entering dormancy under starvation
These responses often involve regulatory networks computing thresholds.
For instance:
if nutrient level < threshold activate starvation response
This is not symbolic reasoning, but it is conditional state-dependent behavior.
4. The Minimum Complexity of Life
One reason minimal cells are fascinating is that they may be close to the lowest complexity capable of sustaining life.
The synthetic organism JCVI-syn3A has roughly:
~500 genes
Yet it can still:
- metabolize nutrients
- repair cellular components
- regulate internal chemistry
- replicate itself
That means the basic control architecture of life may exist at surprisingly small scale.
5. Life as an Active Inference System
Some theoretical neuroscientists interpret living organisms through the lens of predictive control.
One influential idea is the **Free Energy Principle proposed by Karl Friston.
Very roughly:
Living systems maintain themselves by minimizing surprise relative to their environment.
In practice this means:
- sensing environmental states
- adjusting internal processes
- acting to maintain viability
Even bacteria perform behaviors that resemble primitive homeostatic inference.
6. Why Minimal Cells Matter for AI
If the smallest living systems already contain the building blocks of intelligence, then understanding them may reveal the minimal architecture required for adaptive behavior.
Key ingredients include:
- feedback control
- energy management
- information storage
- self-repair
- reproduction
These features appear repeatedly in complex intelligent systems.
7. Digital Cells as Experimental Platforms
With a full simulation of a minimal cell, researchers can now explore questions like:
- What regulatory networks are essential for survival?
- How does information propagate through biochemical systems?
- Which feedback loops stabilize cellular behavior?
These experiments are extremely difficult in real organisms but relatively easy in simulation.
That makes digital cells powerful tools for studying the origins of adaptive systems.
8. The Surprising Connection to AGI
The deepest implication is that life and intelligence may share a common systems architecture.
Both involve:
| Feature | Biological system | Intelligent system |
|---|---|---|
| state sensing | molecular receptors | sensors / inputs |
| internal modeling | regulatory networks | world models |
| adaptive response | metabolic control | planning / action |
| persistence | homeostasis | goal maintenance |
Understanding the minimal version of this architecture in cells could inform future AI system design.
✅ In essence
Minimal cells may represent the smallest naturally occurring systems that maintain themselves through information-driven feedback control.
That is why some complexity scientists think they exhibit a form of proto-intelligence.
The UIUC digital simulation of JCVI-syn3A may therefore allow scientists to study the boundary between chemistry, life, and intelligence in a way that was previously impossible.
If you’re interested, there is an even more provocative implication emerging from this line of thinking:
Some researchers suspect that intelligence might ultimately be understood as a special case of the same principles that govern living systems.
That idea could have very large consequences for both AI and biology.
W.H.L.: Please share the idea.
ChatGPT: The provocative idea some researchers are exploring is that intelligence may not be a separate phenomenon from life, but rather a continuation of the same organizing principles that make living systems possible. In other words, intelligence might emerge naturally once systems reach a certain level of self-maintaining complexity.
This perspective draws from systems biology, cybernetics, and complexity science, and the new digital simulation of the minimal organism JCVI-syn3A at University of Illinois Urbana–Champaign gives scientists a rare platform to examine those principles in detail.
Below is the core logic of this idea.
1. Life Is a Self-Maintaining Information System
All living systems share a defining property:
they actively maintain their own organization.
This requires constant regulation of:
- energy flows
- molecular composition
- internal structure
The theory of autopoiesis developed by Humberto Maturana and Francisco Varela describes living organisms as networks that continuously regenerate themselves.
To do this, cells must:
- sense their environment
- process information
- respond adaptively
Those are the same functional elements associated with intelligence.
2. Intelligence Extends the Same Feedback Principles
From a systems perspective, both living organisms and intelligent systems rely on feedback loops.
For example:
| System | Feedback function |
|---|---|
| bacterial cell | regulate metabolism |
| nervous system | regulate behavior |
| AI agent | regulate action policies |
Cybernetics pioneer Norbert Wiener argued that feedback control is the fundamental mechanism behind both biological regulation and intelligent behavior.
In this view, intelligence is simply feedback control operating at larger scales and higher abstraction levels.
3. Increasing Layers of Adaptive Control
Complexity scientists sometimes describe life and intelligence as a hierarchy of adaptive systems.
A simplified progression might look like:
chemical self-organization ↓minimal life (cells) ↓multicellular organisms ↓nervous systems ↓cognitive intelligence
Each stage adds:
- more sensors
- more internal modeling
- more sophisticated control mechanisms
But the underlying principles remain similar.
4. Prediction as a Core Principle
Many modern theories of intelligence emphasize prediction.
Organisms constantly anticipate environmental changes in order to survive.
The **Free Energy Principle proposed by Karl Friston frames biological systems as entities that minimize surprise by predicting their environment.
Examples:
- bacteria moving toward nutrients
- animals predicting predator behavior
- humans planning future actions
These behaviors can be interpreted as increasingly sophisticated predictive models of the world.
5. Cells Already Exhibit Proto-Prediction
Even simple cells perform primitive forms of predictive control.
For instance:
- bacteria can anticipate nutrient gradients
- regulatory networks prepare stress responses
- metabolic systems adjust before energy depletion occurs
These responses often involve time-dependent regulatory circuits, meaning the system is not merely reacting—it is anticipating likely future states.
6. Intelligence as a Phase Transition
Some complexity researchers propose that intelligence emerges when adaptive systems cross a certain threshold.
Below that threshold:
- systems regulate internal chemistry
Above it:
- systems build internal models of the external world
This transition might resemble a phase change, similar to how liquid water becomes ice when temperature drops.
Minimal cells could be close to that boundary.
7. Why Whole-Cell Simulations Matter
The simulation of JCVI-syn3A gives scientists a tool to explore questions like:
- Which feedback loops stabilize life?
- How do regulatory networks maintain order?
- What minimal structures enable adaptive behavior?
Because the system is fully simulated, researchers can run controlled experiments on the architecture of life itself.
8. Implications for AI
If intelligence and life share the same underlying principles, future AI systems might benefit from architectures that incorporate features common in living systems:
- self-maintenance
- energy-aware computation
- distributed control
- adaptive feedback loops
These ideas already appear in fields like:
- embodied AI
- adaptive robotics
- neuromorphic computing
Understanding biological organization could therefore guide the next generation of AI architectures.
9. A Unified View of Intelligence
From this perspective, intelligence is not a mysterious property that suddenly appears in brains or machines.
Instead, it may represent the upper end of a continuum of adaptive complexity that begins with simple living cells.
The chain might look like:
chemistry → life → adaptive systems → intelligence
Each stage builds on the previous one.
✅ The deeper implication
The simulation of minimal cells like JCVI-syn3A might allow scientists to study the earliest stages of that continuum, where chemistry becomes life and adaptive regulation begins to resemble intelligence.
Understanding that transition could reshape both biology and artificial intelligence.
If you’re interested, there is an even more speculative idea emerging from this line of thinking:
some researchers suspect that intelligence may ultimately be constrained by the same thermodynamic laws that govern living systems.
That connection between information, energy, and intelligence is becoming an active area of research.
W.H.L.: There is a lot more to talk about, but we have to wrap up for now. Thanks to everyone for today’s conversation. See you all soon. Bye!

Leave a comment